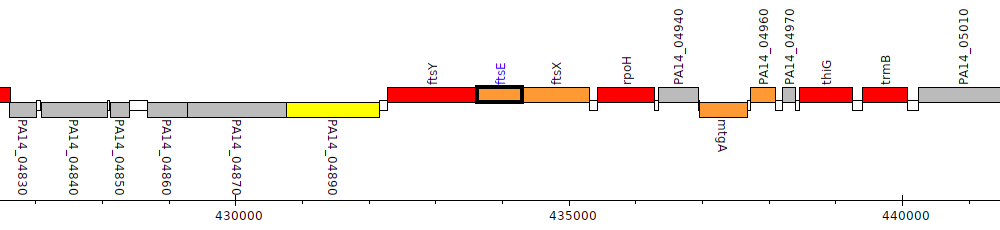
Pseudomonas aeruginosa UCBPP-PA14, PA14_04910 (ftsE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00005
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0051301 | cell division |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02673
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau02010 | ABC transporters | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 222 | 4.8E-66 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 27 | 214 | 6.8E-12 |
| PANTHER | PTHR24220 | IMPORT ATP-BINDING PROTEIN | IPR015854 | ABC transporter, lipoprotein release, LolD-like | 1 | 221 | 7.4E-60 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 218 | 1.06E-66 |
| Pfam | PF00005 | ABC transporter | IPR003439 | ABC transporter-like, ATP-binding domain | 18 | 165 | 1.2E-34 |
| FunFam | G3DSA:3.40.50.300:FF:000056 | Cell division ATP-binding protein FtsE | - | - | 1 | 220 | 9.1E-87 |
| NCBIfam | TIGR02673 | JCVI: cell division ATP-binding protein FtsE | IPR005286 | Cell division protein FtsE | 1 | 214 | 3.5E-85 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.