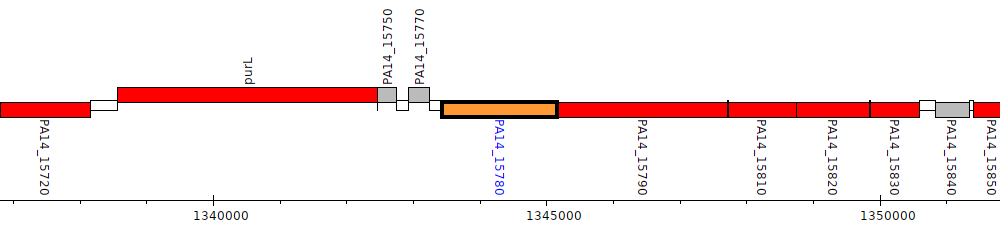
Pseudomonas aeruginosa UCBPP-PA14, PA14_15780
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0019866 | organelle inner membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01998
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02378
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015572 | N-acetylglucosamine transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01998
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009401 | phosphoenolpyruvate-dependent sugar phosphotransferase system |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1360.60
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008982 | protein-N(PI)-phosphohistidine-sugar phosphotransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1360.60
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau02060 | Phosphotransferase system (PTS) | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR01998 | JCVI: N-acetylglucosamine-specific PTS transporter subunit IIBC | IPR010974 | Phosphotransferase system, N-acetylglucosamine-specific IIBC component | 7 | 457 | 0.0 |
| SUPERFAMILY | SSF55604 | Glucose permease domain IIB | IPR036878 | Glucose permease domain IIB | 494 | 567 | 1.31E-8 |
| SUPERFAMILY | SSF55604 | Glucose permease domain IIB | IPR036878 | Glucose permease domain IIB | 385 | 459 | 3.4E-16 |
| Pfam | PF00367 | phosphotransferase system, EIIB | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 389 | 419 | 4.7E-9 |
| Pfam | PF02378 | Phosphotransferase system, EIIC | IPR003352 | Phosphotransferase system, EIIC | 14 | 303 | 3.0E-72 |
| Gene3D | G3DSA:3.30.1360.60 | Glucose permease domain IIB | IPR036878 | Glucose permease domain IIB | 381 | 461 | 4.5E-22 |
| Gene3D | G3DSA:3.30.1360.60 | Glucose permease domain IIB | IPR036878 | Glucose permease domain IIB | 490 | 569 | 6.0E-12 |
| CDD | cd00212 | PTS_IIB_glc | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 384 | 457 | 5.18673E-20 |
| NCBIfam | TIGR00826 | JCVI: glucose PTS transporter subunit EIIB | IPR001996 | Phosphotransferase system, IIB component, type 1 | 360 | 444 | 4.5E-14 |
| PANTHER | PTHR30009 | CYTOCHROME C-TYPE SYNTHESIS PROTEIN AND PTS TRANSMEMBRANE COMPONENT | - | - | 5 | 461 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.