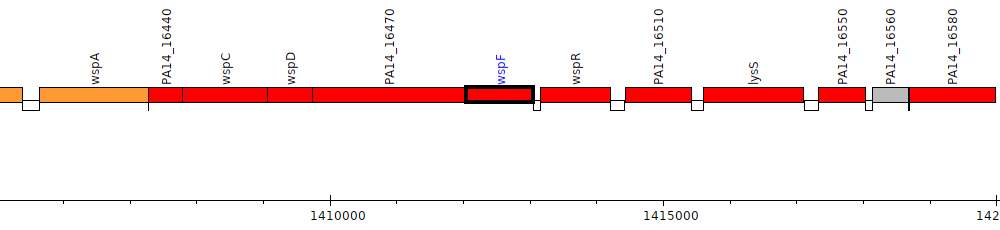
Pseudomonas aeruginosa UCBPP-PA14, PA14_16480 (wspF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008984 | protein-glutamate methylesterase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000156 | phosphorelay response regulator activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000876
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52172 | CheY-like | IPR011006 | CheY-like superfamily | 1 | 104 | 9.53E-21 |
| PANTHER | PTHR42872 | PROTEIN-GLUTAMATE METHYLESTERASE/PROTEIN-GLUTAMINE GLUTAMINASE | - | - | 1 | 334 | 2.7E-100 |
| Gene3D | G3DSA:3.40.50.2300 | - | - | - | 1 | 129 | 1.0E-21 |
| SUPERFAMILY | SSF52738 | Methylesterase CheB, C-terminal domain | IPR035909 | Methylesterase CheB, C-terminal | 151 | 334 | 1.96E-61 |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 4 | 102 | 3.3E-16 |
| SMART | SM00448 | REC_2 | IPR001789 | Signal transduction response regulator, receiver domain | 1 | 118 | 6.3E-10 |
| Pfam | PF01339 | CheB methylesterase | IPR000673 | Signal transduction response regulator, chemotaxis, protein-glutamate methylesterase | 154 | 330 | 5.6E-57 |
| CDD | cd16432 | CheB_Rec | IPR000673 | Signal transduction response regulator, chemotaxis, protein-glutamate methylesterase | 153 | 332 | 3.29997E-78 |
| CDD | cd17541 | REC_CheB-like | - | - | 1 | 124 | 3.33424E-48 |
| PIRSF | PIRSF000876 | RR_chemtxs_CheB | IPR008248 | Protein-glutamate methylesterase/protein-glutamine glutaminase, CheB type | 1 | 335 | 5.2E-98 |
| Gene3D | G3DSA:3.40.50.180 | - | IPR035909 | Methylesterase CheB, C-terminal | 146 | 335 | 1.4E-65 |
| Hamap | MF_00099 | Protein-glutamate methylesterase/protein-glutamine glutaminase [cheB]. | IPR008248 | Protein-glutamate methylesterase/protein-glutamine glutaminase, CheB type | 1 | 335 | 32.583981 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.