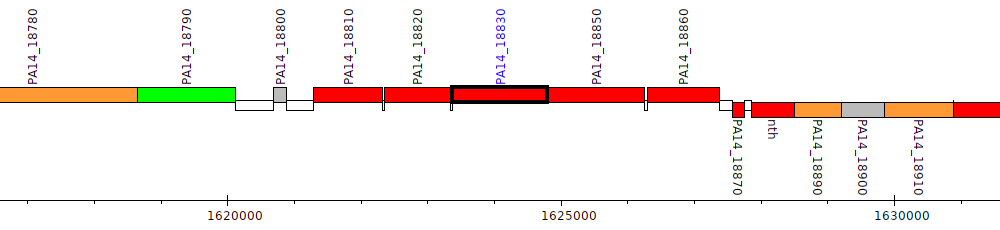
Pseudomonas aeruginosa UCBPP-PA14, PA14_18830
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004018 | N6-(1,2-dicarboxyethyl)AMP AMP-lyase (fumarate-forming) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00928
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009152 | purine ribonucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00928
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48557
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | inosine-5'-phosphate biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | inosine-5'-phosphate biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau00250 | Alanine, aspartate and glutamate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | inosine-5'-phosphate biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosine ribonucleotides <i>de novo</i> biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43172 | ADENYLOSUCCINATE LYASE | - | - | 2 | 450 | 5.7E-124 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 102 | 120 | 8.6E-24 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 447 | 477 | - |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 101 | 123 | 2.4E-10 |
| SMART | SM00998 | ADSL_C_2 | IPR019468 | Adenylosuccinate lyase C-terminal | 366 | 445 | 2.4E-25 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 147 | 165 | 8.6E-24 |
| Pfam | PF00206 | Lyase | IPR022761 | Fumarate lyase, N-terminal | 17 | 298 | 7.6E-42 |
| Gene3D | G3DSA:1.20.200.10 | Fumarase/aspartase (Central domain) | - | - | 3 | 363 | 1.6E-104 |
| Gene3D | G3DSA:1.10.40.30 | - | - | - | 366 | 450 | 1.3E-21 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 273 | 287 | - |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 277 | 293 | 2.4E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 457 | 471 | - |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 277 | 293 | 8.6E-24 |
| SUPERFAMILY | SSF48557 | L-aspartase-like | IPR008948 | L-Aspartase-like | 11 | 445 | 1.4E-115 |
| Pfam | PF10397 | Adenylosuccinate lyase C-terminus | IPR019468 | Adenylosuccinate lyase C-terminal | 367 | 442 | 1.1E-15 |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 142 | 162 | 2.4E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 268 | 287 | - |
| NCBIfam | TIGR00928 | JCVI: adenylosuccinate lyase | IPR004769 | Adenylosuccinate lyase | 15 | 444 | 9.4E-99 |
| CDD | cd01597 | pCLME | - | - | 10 | 447 | 0.0 |
| PRINTS | PR00149 | Fumarate lyase superfamily signature | IPR000362 | Fumarate lyase family | 232 | 259 | 8.6E-24 |
| PRINTS | PR00145 | Argininosuccinate lyase family signature | - | - | 232 | 256 | 2.4E-10 |
| FunFam | G3DSA:1.20.200.10:FF:000014 | 3-carboxy-cis,cis-muconate cycloisomerase | - | - | 3 | 363 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.