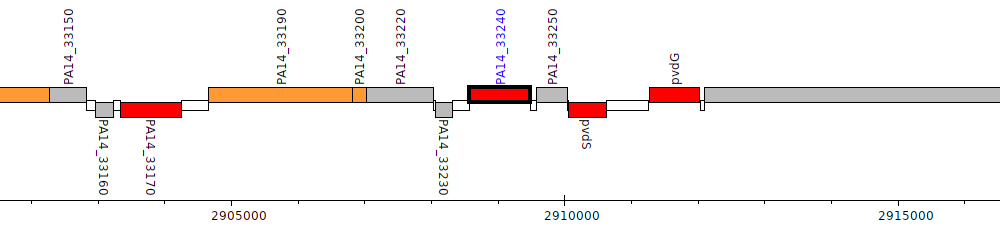
Pseudomonas aeruginosa UCBPP-PA14, PA14_33240
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008976 | polyphosphate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03707
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006793 | phosphorus metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03707
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR34383 | POLYPHOSPHATE:AMP PHOSPHOTRANSFERASE-RELATED | - | - | 21 | 293 | 1.4E-115 |
| PIRSF | PIRSF028756 | PPK2 | IPR016898 | Polyphosphate phosphotransferase | 12 | 304 | 0.0 |
| Pfam | PF03976 | Polyphosphate kinase 2 (PPK2) | IPR022488 | Polyphosphate kinase-2-related | 49 | 274 | 1.7E-115 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 80 | 268 | 3.32E-32 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 14 | 288 | 1.6E-99 |
| NCBIfam | TIGR03707 | JCVI: polyphosphate kinase 2 | IPR022486 | Polyphosphate kinase 2, PA0141 | 49 | 274 | 1.5E-115 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.