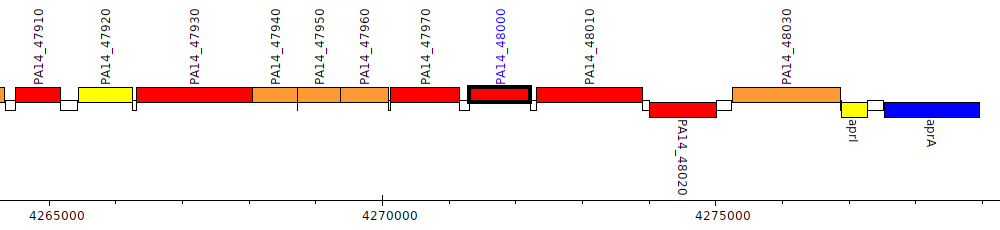
Pseudomonas aeruginosa UCBPP-PA14, PA14_48000
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016829 | lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR12128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.20.20.70:FF:000281 | Probable dihydrodipicolinate synthetase | - | - | 4 | 295 | 0.0 |
| SUPERFAMILY | SSF51569 | Aldolase | - | - | 3 | 292 | 6.38E-80 |
| PIRSF | PIRSF001365 | DHDPS | IPR002220 | DapA-like | 3 | 301 | 2.8E-75 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 4 | 295 | 4.2E-80 |
| SMART | SM01130 | DHDPS_2 | IPR002220 | DapA-like | 6 | 294 | 7.1E-86 |
| PANTHER | PTHR12128 | DIHYDRODIPICOLINATE SYNTHASE | IPR002220 | DapA-like | 4 | 293 | 8.7E-51 |
| CDD | cd00408 | DHDPS-like | - | - | 10 | 290 | 1.09031E-92 |
| Pfam | PF00701 | Dihydrodipicolinate synthetase family | IPR002220 | DapA-like | 8 | 291 | 6.7E-59 |
| PRINTS | PR00146 | Dihydrodipicolinate synthase signature | IPR002220 | DapA-like | 76 | 94 | 1.2E-9 |
| PRINTS | PR00146 | Dihydrodipicolinate synthase signature | IPR002220 | DapA-like | 133 | 150 | 1.2E-9 |
| PRINTS | PR00146 | Dihydrodipicolinate synthase signature | IPR002220 | DapA-like | 40 | 61 | 1.2E-9 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.