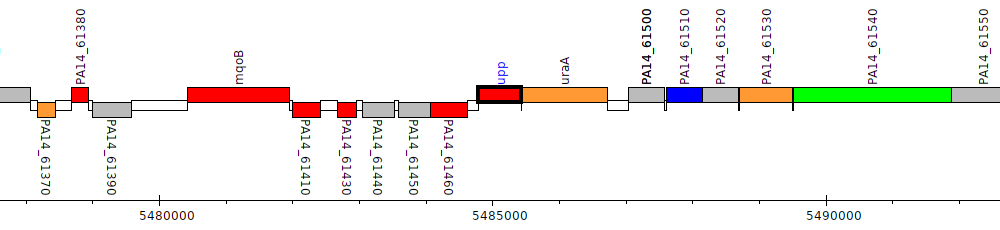
Pseudomonas aeruginosa UCBPP-PA14, PA14_61470 (upp)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006223 | uracil salvage |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01091
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004845 | uracil phosphoribosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01091
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_01218_B | Uracil phosphoribosyltransferase [upp]. | IPR034332 | Uracil phosphoribosyltransferase, bacterial-type | 2 | 208 | 44.828754 |
| Gene3D | G3DSA:3.40.50.2020 | - | IPR029057 | Phosphoribosyltransferase-like | 1 | 208 | 1.0E-71 |
| SUPERFAMILY | SSF53271 | PRTase-like | IPR029057 | Phosphoribosyltransferase-like | 3 | 206 | 4.33E-65 |
| FunFam | G3DSA:3.40.50.2020:FF:000003 | Uracil phosphoribosyltransferase | - | - | 1 | 208 | 7.1E-90 |
| NCBIfam | TIGR01091 | JCVI: uracil phosphoribosyltransferase | IPR005765 | Uracil phosphoribosyl transferase | 3 | 208 | 8.6E-88 |
| PANTHER | PTHR32315 | ADENINE PHOSPHORIBOSYLTRANSFERASE | - | - | 4 | 181 | 7.6E-20 |
| CDD | cd06223 | PRTases_typeI | IPR000836 | Phosphoribosyltransferase domain | 68 | 182 | 2.88343E-18 |
| Pfam | PF14681 | Uracil phosphoribosyltransferase | - | - | 7 | 207 | 2.9E-63 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.