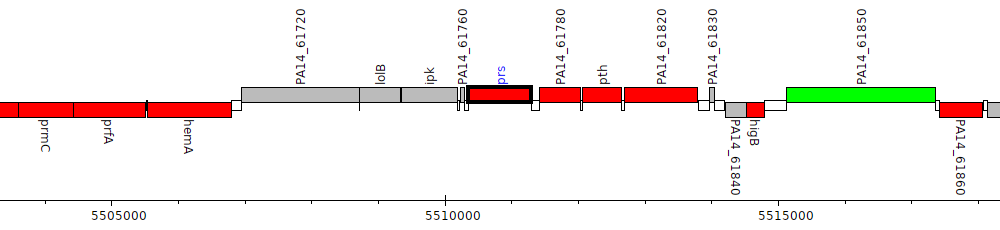
Pseudomonas aeruginosa UCBPP-PA14, PA14_61770 (prs)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004749 | ribose phosphate diphosphokinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF14572
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009165 | nucleotide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF14572
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF14572
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06223 | PRTases_typeI | IPR000836 | Phosphoribosyltransferase domain | 149 | 273 | 7.19707E-27 |
| FunFam | G3DSA:3.40.50.2020:FF:000001 | Ribose-phosphate pyrophosphokinase | - | - | 7 | 160 | 3.7E-70 |
| Gene3D | G3DSA:3.40.50.2020 | - | IPR029057 | Phosphoribosyltransferase-like | 148 | 289 | 0.0 |
| SMART | SM01400 | Pribosyltran_N_2 | - | - | 4 | 121 | 1.0E-76 |
| Hamap | MF_00583_B | Putative ribose-phosphate pyrophosphokinase [prs]. | IPR037515 | Ribose-phosphate pyrophosphokinase, bacterial-type | 4 | 313 | 48.28606 |
| NCBIfam | TIGR01251 | JCVI: ribose-phosphate diphosphokinase | IPR005946 | Ribose-phosphate pyrophosphokinase | 4 | 313 | 2.7E-123 |
| PANTHER | PTHR10210 | RIBOSE-PHOSPHATE DIPHOSPHOKINASE FAMILY MEMBER | IPR005946 | Ribose-phosphate pyrophosphokinase | 1 | 313 | 2.2E-125 |
| Pfam | PF13793 | N-terminal domain of ribose phosphate pyrophosphokinase | IPR029099 | Ribose-phosphate pyrophosphokinase, N-terminal domain | 4 | 121 | 1.2E-48 |
| Gene3D | G3DSA:3.40.50.2020 | - | IPR029057 | Phosphoribosyltransferase-like | 7 | 304 | 0.0 |
| SUPERFAMILY | SSF53271 | PRTase-like | IPR029057 | Phosphoribosyltransferase-like | 69 | 305 | 6.07E-64 |
| Pfam | PF14572 | Phosphoribosyl synthetase-associated domain | IPR005946 | Ribose-phosphate pyrophosphokinase | 201 | 313 | 3.7E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.