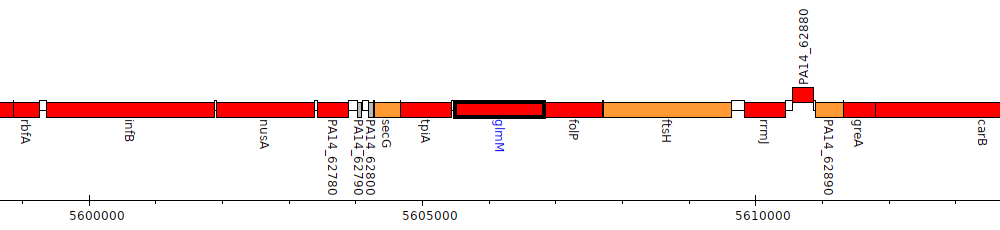
Pseudomonas aeruginosa UCBPP-PA14, PA14_62840 (glmM)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008966 | phosphoglucosamine mutase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4749
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10913078 | |
| Molecular Function | GO:0004614 | phosphoglucomutase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4749
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10913078 | |
| Molecular Function | GO:0004615 | phosphomannomutase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4749
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10913078 | |
| Biological Process | GO:0009252 | peptidoglycan biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4749
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10913078 | |
| Molecular Function | GO:0008966 | phosphoglucosamine mutase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01455
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53738
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016868 | intramolecular transferase activity, phosphotransferases |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53738
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01455
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0071704 | organic substance metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55957
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00509 | Phosphoglucomutase/phosphomannomutase family signature | IPR005841 | Alpha-D-phosphohexomutase superfamily | 94 | 108 | 9.2E-15 |
| Gene3D | G3DSA:3.30.310.50 | - | - | - | 369 | 443 | 7.8E-21 |
| Pfam | PF00408 | Phosphoglucomutase/phosphomannomutase, C-terminal domain | IPR005843 | Alpha-D-phosphohexomutase, C-terminal | 372 | 440 | 2.4E-11 |
| Hamap | MF_01554_B | Phosphoglucosamine mutase [glmM]. | IPR006352 | Phosphoglucosamine mutase, bacterial type | 2 | 443 | 46.768997 |
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 173 | 360 | 9.6E-66 |
| Pfam | PF02880 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain III | IPR005846 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain III | 257 | 362 | 1.8E-30 |
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 2 | 153 | 1.1E-41 |
| CDD | cd05802 | GlmM | IPR006352 | Phosphoglucosamine mutase, bacterial type | 5 | 437 | 0.0 |
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 256 | 338 | 9.6E-66 |
| PANTHER | PTHR42946 | PHOSPHOHEXOSE MUTASE | - | - | 2 | 444 | 7.4E-90 |
| FunFam | G3DSA:3.40.120.10:FF:000001 | Phosphoglucosamine mutase | - | - | 2 | 153 | 5.9E-54 |
| FunFam | G3DSA:3.30.310.50:FF:000001 | Phosphoglucosamine mutase | - | - | 369 | 444 | 5.4E-27 |
| PRINTS | PR00509 | Phosphoglucomutase/phosphomannomutase family signature | IPR005841 | Alpha-D-phosphohexomutase superfamily | 233 | 248 | 9.2E-15 |
| SUPERFAMILY | SSF53738 | Phosphoglucomutase, first 3 domains | IPR016055 | Alpha-D-phosphohexomutase, alpha/beta/alpha I/II/III | 2 | 180 | 9.81E-47 |
| SUPERFAMILY | SSF55957 | Phosphoglucomutase, C-terminal domain | IPR036900 | Alpha-D-phosphohexomutase, C-terminal domain superfamily | 357 | 444 | 1.44E-20 |
| SUPERFAMILY | SSF53738 | Phosphoglucomutase, first 3 domains | IPR016055 | Alpha-D-phosphohexomutase, alpha/beta/alpha I/II/III | 155 | 256 | 2.43E-31 |
| Pfam | PF02878 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain I | IPR005844 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain I | 3 | 134 | 4.2E-44 |
| NCBIfam | TIGR01455 | JCVI: phosphoglucosamine mutase | IPR006352 | Phosphoglucosamine mutase, bacterial type | 6 | 441 | 0.0 |
| SUPERFAMILY | SSF53738 | Phosphoglucomutase, first 3 domains | IPR016055 | Alpha-D-phosphohexomutase, alpha/beta/alpha I/II/III | 257 | 363 | 1.18E-29 |
| Pfam | PF02879 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain II | IPR005845 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain II | 156 | 253 | 6.2E-18 |
| PRINTS | PR00509 | Phosphoglucomutase/phosphomannomutase family signature | IPR005841 | Alpha-D-phosphohexomutase superfamily | 173 | 192 | 9.2E-15 |
| FunFam | G3DSA:3.40.120.10:FF:000003 | Phosphoglucosamine mutase | - | - | 172 | 274 | 1.5E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.