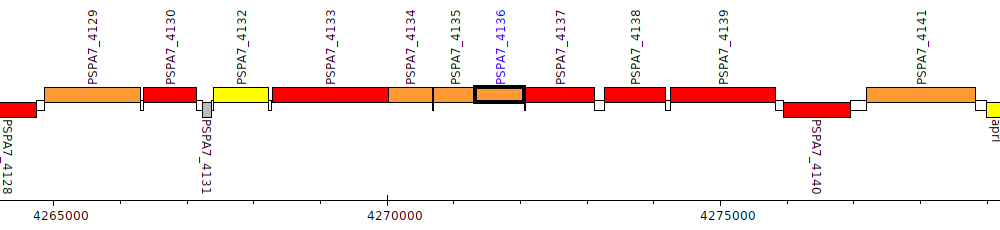
Pseudomonas aeruginosa PA7, PSPA7_4136
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0015424 | ATPase-coupled amino acid transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF039085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0003333 | amino acid transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF039085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00005
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF039085 | ABC_ATPase_HisP | IPR030679 | ABC-type amino acid transport system, ATPase component, HisP-type | 1 | 240 | 2.5E-104 |
| Pfam | PF00005 | ABC transporter | IPR003439 | ABC transporter-like, ATP-binding domain | 18 | 165 | 9.0E-34 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 235 | 3.64E-75 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 240 | 7.4E-73 |
| SMART | SM00382 | AAA_5 | IPR003593 | AAA+ ATPase domain | 26 | 213 | 1.0E-15 |
| FunFam | G3DSA:3.40.50.300:FF:000020 | Amino acid ABC transporter ATP-binding component | - | - | 1 | 240 | 8.7E-105 |
| PANTHER | PTHR43166 | AMINO ACID IMPORT ATP-BINDING PROTEIN | - | - | 1 | 238 | 3.0E-81 |
| CDD | cd03262 | ABC_HisP_GlnQ | - | - | 2 | 212 | 1.13655E-123 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.