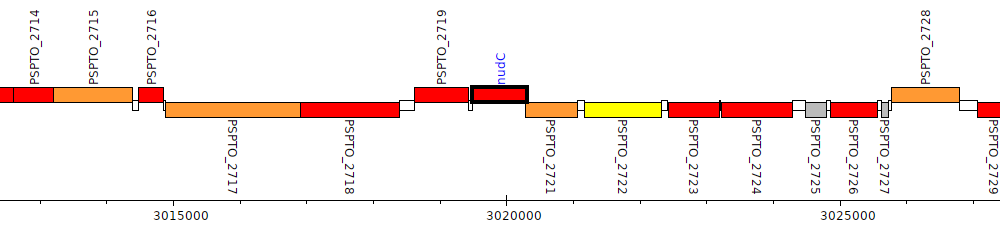
Pseudomonas syringae pv. tomato DC3000, PSPTO_2720 (nudC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0045454 | cell redox homeostasis | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
24989777 | Reviewed by curator |
| Molecular Function | GO:0016787 | hydrolase activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
24989777 | Reviewed by curator |
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF09297
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000210 | NAD+ diphosphatase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00297
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF09297
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pst00760 | Nicotinate and nicotinamide metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pst01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR42904 | NUDIX HYDROLASE, NUDC SUBFAMILY | - | - | 79 | 268 | 6.2E-53 |
| Pfam | PF09297 | NADH pyrophosphatase zinc ribbon domain | IPR015376 | Zinc ribbon, NADH pyrophosphatase | 109 | 140 | 1.3E-10 |
| CDD | cd03429 | NADH_pyrophosphatase | - | - | 145 | 267 | 1.48057E-69 |
| Pfam | PF00293 | NUDIX domain | IPR000086 | NUDIX hydrolase domain | 146 | 257 | 7.9E-19 |
| Hamap | MF_00297 | NAD-capped RNA hydrolase NudC [nudC]. | IPR022925 | NAD-capped RNA hydrolase NudC | 8 | 273 | 49.640221 |
| Gene3D | G3DSA:3.90.79.10 | Nucleoside Triphosphate Pyrophosphohydrolase | - | - | 142 | 271 | 4.0E-39 |
| Gene3D | G3DSA:3.90.79.20 | - | - | - | 4 | 141 | 3.0E-25 |
| SUPERFAMILY | SSF55811 | Nudix | IPR015797 | NUDIX hydrolase-like domain superfamily | 95 | 261 | 1.12E-42 |
| Pfam | PF09296 | NADH pyrophosphatase-like rudimentary NUDIX domain | IPR015375 | NADH pyrophosphatase-like, N-terminal | 23 | 107 | 2.6E-5 |
| SUPERFAMILY | SSF55811 | Nudix | IPR015797 | NUDIX hydrolase-like domain superfamily | 7 | 112 | 3.12E-22 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.