Pseudomonas aeruginosa PAO1, PA0423 (pasP)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
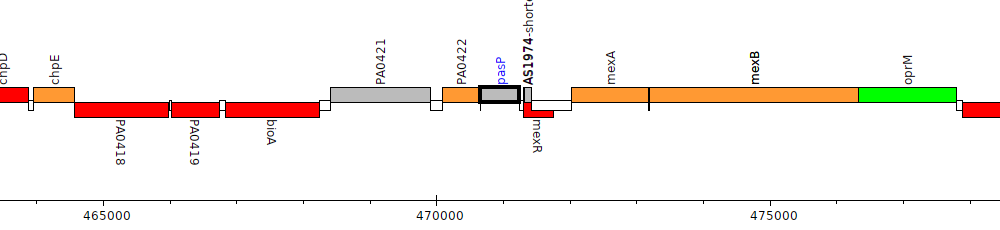
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA0423
|
| Name |
pasP
Synonym: pasP Synonym: yceI |
| Replicon | chromosome |
| Genomic location | 470662 - 471237 (+ strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_249114.1 |
| GI | 15595620 |
| Affymetrix | PA0423_at |
| Entrez | 877902 |
| GenBank | AAG03812.1 |
| INSDC | AAG03812.1 |
| NCBI Locus Tag | PA0423 |
| protein_id(GenBank) | gb|AAG03812.1|AE004479_7|gnl|PseudoCAP|PA0423 |
| TIGR | NTL03PA00424 |
| UniParc | UPI00000C504B |
| UniProtKB Acc | Q9I690 |
| UniProtKB ID | Y423_PSEAE |
| UniRef100 | UniRef100_B7V411 |
| UniRef50 | UniRef50_Q32E67 |
| UniRef90 | UniRef90_B7V411 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
PasP
|
| Synonyms |
PasP |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
Detected by MALDI-TOF/TOF (PMID:19333994).
Detected using LC-MS/MS (20.78 kDa, PMID:21751344)
Verified using two-dimensional gel electrophoresis, MALDI-TOF MS and peptide mass mapping(pI:6.1, molecular weight:20.8, PMID:21674802)
|
| Charge (pH 7) | -1.04 |
| Kyte-Doolittle Hydrophobicity Value | -0.380 |
| Molecular Weight (kDa) | 20.8 |
| Isoelectric Point (pI) | 6.53 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7BWL | ANTIBIOTIC | 04/14/20 | Structure of antibiotic sequester from Pseudomonas aerurinosa | Pseudomonas aeruginosa PAO1 | 2.25 | X-RAY DIFFRACTION | 100.0 |
Virulence Evidence
| Source 1 | |
| Database | PseudoCAP |
| Category | Pseudomonas aeruginosa small protease (PASP) |
| Evidence |
ECO:0000314
direct assay evidence used in manual assertion
|
| Host Organism | Oryctolagus cuniculus |
| Notes | |
| Infection model | P. aeruginosa corneal infection of New Zealand white rabbit |
| Reference | 16186360 |
| Source 2 | |
| Database | PseudoCAP |
| Category | Pseudomonas aeruginosa small protease (PASP) |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Mus musculus |
| Notes | |
| Infection model | Mouse scratch model of P. aeruginosa keratitis |
| Reference | 23548618 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 310 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000414 (536 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA0423
Search term: pasP
Search term: PasP
|
Human Homologs
References
|
Identification of a novel secreted protease from Pseudomonas aeruginosa that causes corneal erosions.
Marquart ME, Caballero AR, Chomnawang M, Thibodeaux BA, Twining SS, O'Callaghan RJ
Invest Ophthalmol Vis Sci 2005 Oct;46(10):3761-8
PubMed ID: 16186360
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|
|
Proteomics of the oxidative stress response induced by hydrogen peroxide and paraquat reveals a novel AhpC-like protein in Pseudomonas aeruginosa.
Hare NJ, Scott NE, Shin EH, Connolly AM, Larsen MR, Palmisano G, Cordwell SJ
Proteomics 2011 Aug;11(15):3056-69
PubMed ID: 21674802
|
|
Proteomic analysis of outer membrane vesicles derived from Pseudomonas aeruginosa.
Choi DS, Kim DK, Choi SJ, Lee J, Choi JP, Rho S, Park SH, Kim YK, Hwang D, Gho YS
Proteomics 2011 Aug;11(16):3424-9
PubMed ID: 21751344
|
|
Mep72, a metzincin protease that is preferentially secreted by biofilms of Pseudomonas aeruginosa.
Passmore IJ, Nishikawa K, Lilley KS, Bowden SD, Chung JC, Welch M
J Bacteriol 2015 Feb 15;197(4):762-73
PubMed ID: 25488299
|