Pseudomonas aeruginosa PAO1, PA4109 (ampR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
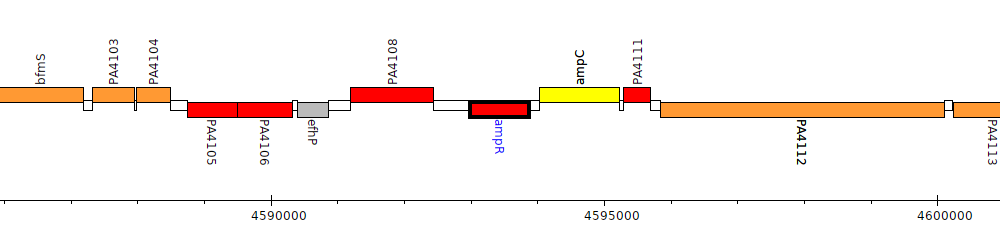
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA4109
|
| Name |
ampR
|
| Replicon | chromosome |
| Genomic location | 4592990 - 4593880 (- strand) |
| Transposon Mutants | 2 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 2 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_252798.1 |
| GI | 15599304 |
| Affymetrix | PA4109_ampR_at |
| Entrez | 877983 |
| GenBank | AAG07496.1 |
| INSDC | CAA38523.2 |
| NCBI Locus Tag | PA4109 |
| protein_id(GenBank) | gb|AAG07496.1|AE004827_5|gnl|PseudoCAP|PA4109 |
| TIGR | NTL03PA04109 |
| UniParc | UPI0000125A26 |
| UniProtKB Acc | P24734 |
| UniProtKB ID | AMPR_PSEAE |
| UniRef100 | UniRef100_P24734 |
| UniRef50 | UniRef50_P24734 |
| UniRef90 | UniRef90_P24734 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
transcriptional regulator AmpR
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | 1.53 |
| Kyte-Doolittle Hydrophobicity Value | 0.127 |
| Molecular Weight (kDa) | 32.6 |
| Isoelectric Point (pI) | 7.36 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 5MMH | TRANSCRIPTION | 12/09/16 | The X-Ray Structure of the Effector Domain of the Transcriptional Regulator AmpR of Pseudomonas aeruginosa | Pseudomonas aeruginosa PAO1 | 2.2 | X-RAY DIFFRACTION | 99.5 |
Virulence Evidence
| Source 1 | |
| Database | PseudoCAP |
| Category | AmpR |
| Evidence |
ECO:0000315
mutant phenotype evidence used in manual assertion
|
| Host Organism | Caenorhabditis elegans |
| Notes | |
| Infection model | C. Elegans fast killing (paralytic) assay |
| Reference | 22479525 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 456 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001142 (1134 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA4109
Search term: ampR
Search term: transcriptional regulator AmpR
|
Human Homologs
References
|
Cloning, sequencing and analysis of the structural gene and regulatory region of the Pseudomonas aeruginosa chromosomal ampC beta-lactamase.
Lodge JM, Minchin SD, Piddock LJ, Busby SJ
Biochem J 1990 Dec 15;272(3):627-31
PubMed ID: 2125210
|
|
Investigation of the Pseudomonas aeruginosa ampR gene and its role at the chromosomal ampC beta-lactamase promoter.
Lodge J, Busby S, Piddock L
FEMS Microbiol Lett 1993 Aug 1;111(2-3):315-20
PubMed ID: 8405939
|
|
The regulatory repertoire of Pseudomonas aeruginosa AmpC ß-lactamase regulator AmpR includes virulence genes.
Balasubramanian D, Schneper L, Merighi M, Smith R, Narasimhan G, Lory S, Mathee K
PLoS One 2012;7(3):e34067
PubMed ID: 22479525
|
|
Structural and functional characterization of Pseudomonas aeruginosa global regulator AmpR.
Caille O, Zincke D, Merighi M, Balasubramanian D, Kumari H, Kong KF, Silva-Herzog E, Narasimhan G, Schneper L, Lory S, Mathee K
J Bacteriol 2014 Nov;196(22):3890-902
PubMed ID: 25182487
|
|
Pseudomonas aeruginosa AmpR: an acute-chronic switch regulator.
Balasubramanian D, Kumari H, Mathee K
Pathog Dis 2015 Mar;73(2):1-14
PubMed ID: 25066236
|
|
Role of Pseudomonas aeruginosa AmpR on β-lactam and non-β-lactam transient cross-resistance upon pre-exposure to subinhibitory concentrations of antibiotics.
Kumari H, Balasubramanian D, Zincke D, Mathee K
J. Med. Microbiol. 2014 Apr;63(Pt 4):544-55
PubMed ID: 24464693
|
|
LTQ-XL mass spectrometry proteome analysis expands the Pseudomonas aeruginosa AmpR regulon to include cyclic di-GMP phosphodiesterases and phosphoproteins, and identifies novel open reading frames.
Kumari H, Murugapiran SK, Balasubramanian D, Schneper L, Merighi M, Sarracino D, Lory S, Mathee K
J Proteomics 2014 Jan 16;96:328-342
PubMed ID: 24291602
|
|
Deep sequencing analyses expands the Pseudomonas aeruginosa AmpR regulon to include small RNA-mediated regulation of iron acquisition, heat shock and oxidative stress response.
Balasubramanian D, Kumari H, Jaric M, Fernandez M, Turner KH, Dove SL, Narasimhan G, Lory S, Mathee K
Nucleic Acids Res 2014 Jan;42(2):979-98
PubMed ID: 24157832
|
|
A dynamic and intricate regulatory network determines Pseudomonas aeruginosa virulence.
Balasubramanian D, Schneper L, Kumari H, Mathee K
Nucleic Acids Res 2013 Jan 7;41(1):1-20
PubMed ID: 23143271
|