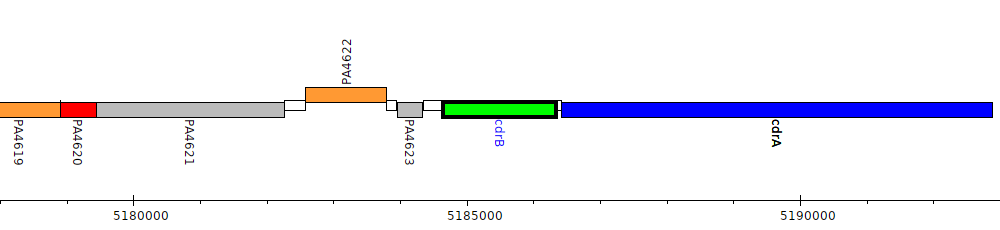
Pseudomonas aeruginosa PAO1, PA4624 (cdrB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA4624
|
| Name |
cdrB
|
| Replicon | chromosome |
| Genomic location | 5184642 - 5186348 (- strand) |
| Transposon Mutants | 12 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 8 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_253314.1 |
| GI | 15599820 |
| Affymetrix | PA4624_at |
| Entrez | 881173 |
| GenBank | AAG08012.1 |
| INSDC | AAG08012.1 |
| NCBI Locus Tag | PA4624 |
| protein_id(GenBank) | gb|AAG08012.1|AE004876_7|gnl|PseudoCAP|PA4624 |
| TIGR | NTL03PA04625 |
| UniParc | UPI00000C5D94 |
| UniProtKB Acc | Q9HVG7 |
| UniProtKB ID | Q9HVG7_PSEAE |
| UniRef100 | UniRef100_Q9HVG7 |
| UniRef50 | UniRef50_Q9HVG7 |
| UniRef90 | UniRef90_Q9HVG7 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
cyclic diguanylate-regulated TPS partner B, CdrB
|
| Synonyms | |
| Evidence for Translation |
Detected by MALDI-TOF/TOF (PMID:19333994).
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -5.83 |
| Kyte-Doolittle Hydrophobicity Value | -0.409 |
| Molecular Weight (kDa) | 63.3 |
| Isoelectric Point (pI) | 5.45 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 152 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000040 (1614 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA4624
Search term: cdrB
Search term: cyclic diguanylate-regulated TPS partner B, CdrB
|
Human Homologs
References
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
Whole-genome random sequencing and assembly of Haemophilus influenzae Rd.
Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Bult CJ, Tomb JF, Dougherty BA, Merrick JM
Science 1995 Jul 28;269(5223):496-512
PubMed ID: 7542800
|
|
Pseudomonas aeruginosa uses a cyclic-di-GMP-regulated adhesin to reinforce the biofilm extracellular matrix.
Borlee BR, Goldman AD, Murakami K, Samudrala R, Wozniak DJ, Parsek MR
Mol Microbiol 2010 Feb;75(4):827-42
PubMed ID: 20088866
|
|
Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, Ascenzi P, Piacentini M, Visca P, Fimia GM
Proteomics 2009 Apr;9(7):1901-15
PubMed ID: 19333994
|