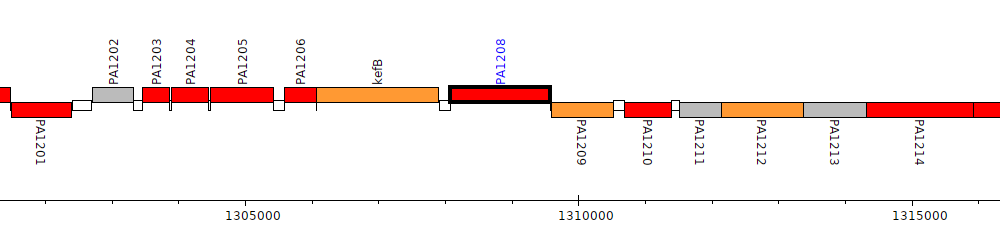
Pseudomonas aeruginosa PAO1, PA1208
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050661 | NADP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00743
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00743
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004499 | N,N-dimethylaniline monooxygenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00743
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 158 | 1.0E-38 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 159 | 252 | 1.0E-11 |
| Pfam | PF00743 | Flavin-binding monooxygenase-like | IPR020946 | Flavin monooxygenase-like | 82 | 217 | 1.1E-8 |
| Pfam | PF13450 | NAD(P)-binding Rossmann-like domain | - | - | 10 | 57 | 2.9E-6 |
| PANTHER | PTHR43872 | MONOOXYGENASE, PUTATIVE (AFU_ORTHOLOGUE AFUA_8G02570)-RELATED | - | - | 3 | 482 | 0.0 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 314 | 446 | 1.4E-6 |
| FunFam | G3DSA:3.50.50.60:FF:000228 | FAD-containing monooxygenase EthA | - | - | 2 | 155 | 3.1E-63 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 4 | 386 | 7.34E-51 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.