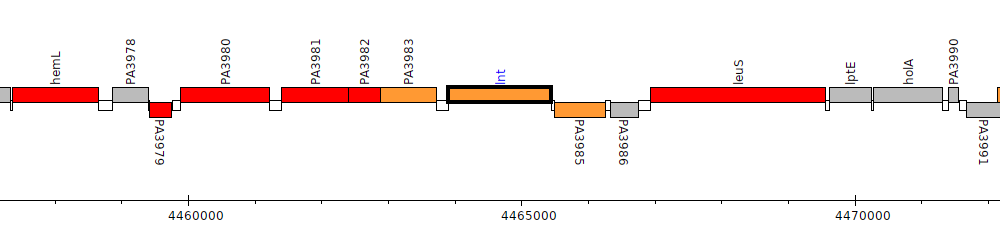
Pseudomonas aeruginosa PAO1, PA3984 (lnt)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006878 | cellular copper ion homeostasis | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8344936 | Reviewed by curator |
| Biological Process | GO:0042160 | lipoprotein modification | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
8344936 | Reviewed by curator |
| Cellular Component | GO:0009276 | Gram-negative-bacterium-type cell wall | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38686
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006807 | nitrogen compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00795
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016410 | N-acyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38686
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042158 | lipoprotein biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR38686
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56317 | Carbon-nitrogen hydrolase | IPR036526 | Carbon-nitrogen hydrolase superfamily | 221 | 484 | 4.19E-47 |
| Hamap | MF_01148 | Apolipoprotein N-acyltransferase [lnt]. | IPR004563 | Apolipoprotein N-acyltransferase | 8 | 489 | 22.896618 |
| NCBIfam | TIGR00546 | JCVI: apolipoprotein N-acyltransferase | IPR004563 | Apolipoprotein N-acyltransferase | 61 | 452 | 7.9E-97 |
| Gene3D | G3DSA:3.60.110.10 | - | IPR036526 | Carbon-nitrogen hydrolase superfamily | 216 | 475 | 7.3E-42 |
| Pfam | PF20154 | Apolipoprotein N-acyltransferase N-terminal domain | IPR045378 | Apolipoprotein N-acyltransferase, N-terminal | 18 | 181 | 8.6E-34 |
| CDD | cd07571 | ALP_N-acyl_transferase | IPR004563 | Apolipoprotein N-acyltransferase | 225 | 484 | 2.51468E-111 |
| Pfam | PF00795 | Carbon-nitrogen hydrolase | IPR003010 | Carbon-nitrogen hydrolase | 230 | 467 | 8.3E-29 |
| PANTHER | PTHR38686 | APOLIPOPROTEIN N-ACYLTRANSFERASE | IPR004563 | Apolipoprotein N-acyltransferase | 2 | 504 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.