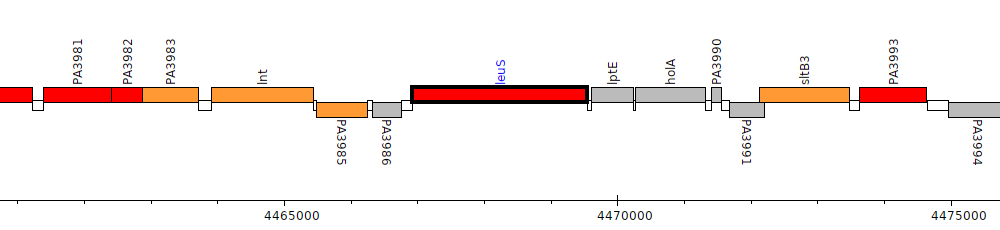
Pseudomonas aeruginosa PAO1, PA3987 (leuS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
10024478 | Reviewed by curator |
| Molecular Function | GO:0004823 | leucine-tRNA ligase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
10024478 | Reviewed by curator |
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0002161 | aminoacyl-tRNA editing activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF13603
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF09334
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004823 | leucine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00049_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00049_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF09334
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006429 | leucyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00049_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00049_B
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | TRNA-CHARGING-PWY | tRNA charging | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Valine, leucine and isoleucine biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Aminoacyl-tRNA biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF13603 | Leucyl-tRNA synthetase, Domain 2 | IPR025709 | Leucyl-tRNA synthetase, editing domain | 221 | 413 | 4.8E-68 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 215 | 234 | 6.8E-51 |
| SUPERFAMILY | SSF47323 | Anticodon-binding domain of a subclass of class I aminoacyl-tRNA synthetases | IPR009080 | Aminoacyl-tRNA synthetase, class Ia, anticodon-binding | 675 | 872 | 9.53E-34 |
| Pfam | PF00133 | tRNA synthetases class I (I, L, M and V) | IPR002300 | Aminoacyl-tRNA synthetase, class Ia | 631 | 666 | 4.9E-7 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 497 | 515 | 6.8E-51 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 153 | 169 | 6.8E-51 |
| FunFam | G3DSA:2.20.28.290:FF:000001 | Leucine--tRNA ligase | - | - | 581 | 637 | 1.6E-24 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 127 | 144 | 6.8E-51 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 410 | 580 | 2.5E-67 |
| Gene3D | G3DSA:2.20.28.290 | - | - | - | 581 | 637 | 1.7E-28 |
| Hamap | MF_00049_B | Leucine--tRNA ligase [leuS]. | IPR002302 | Leucine-tRNA ligase | 4 | 872 | 23.377468 |
| SUPERFAMILY | SSF52374 | Nucleotidylyl transferase | - | - | 5 | 681 | 0.0 |
| Gene3D | G3DSA:3.10.20.590 | - | - | - | 811 | 873 | 5.0E-23 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 536 | 558 | 6.8E-51 |
| Pfam | PF09334 | tRNA synthetases class I (M) | IPR015413 | Methionyl/Leucyl tRNA synthetase | 39 | 171 | 5.9E-19 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 569 | 579 | 6.8E-51 |
| PRINTS | PR00985 | Leucyl-tRNA synthetase signature | IPR002302 | Leucine-tRNA ligase | 185 | 198 | 6.8E-51 |
| FunFam | G3DSA:3.90.740.10:FF:000012 | Leucine--tRNA ligase | - | - | 239 | 419 | 2.3E-85 |
| FunFam | G3DSA:3.10.20.590:FF:000001 | Leucine--tRNA ligase | - | - | 811 | 873 | 1.1E-25 |
| Gene3D | G3DSA:1.10.730.10 | - | - | - | 638 | 810 | 1.4E-63 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 624 | 643 | - |
| NCBIfam | TIGR00396 | JCVI: bacterial-type leucine--tRNA ligase | IPR002302 | Leucine-tRNA ligase | 5 | 872 | 0.0 |
| Gene3D | G3DSA:3.90.740.10 | - | IPR009008 | Valyl/Leucyl/Isoleucyl-tRNA synthetase, editing domain | 236 | 409 | 1.1E-23 |
| FunFam | G3DSA:3.40.50.620:FF:000003 | Leucine--tRNA ligase | - | - | 32 | 252 | 2.9E-111 |
| CDD | cd07958 | Anticodon_Ia_Leu_BEm | - | - | 671 | 789 | 8.32098E-45 |
| Pfam | PF08264 | Anticodon-binding domain of tRNA ligase | IPR013155 | Methionyl/Valyl/Leucyl/Isoleucyl-tRNA synthetase, anticodon-binding | 714 | 834 | 1.3E-14 |
| Gene3D | G3DSA:1.10.730.10 | - | - | - | 6 | 31 | 6.9E-84 |
| PANTHER | PTHR43740 | LEUCYL-TRNA SYNTHETASE | IPR002302 | Leucine-tRNA ligase | 3 | 872 | 0.0 |
| Pfam | PF00133 | tRNA synthetases class I (I, L, M and V) | IPR002300 | Aminoacyl-tRNA synthetase, class Ia | 428 | 584 | 5.5E-8 |
| CDD | cd00812 | LeuRS_core | - | - | 34 | 235 | 1.42592E-87 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 32 | 228 | 6.9E-84 |
| SUPERFAMILY | SSF50677 | ValRS/IleRS/LeuRS editing domain | IPR009008 | Valyl/Leucyl/Isoleucyl-tRNA synthetase, editing domain | 226 | 424 | 1.26E-49 |
| FunFam | G3DSA:1.10.730.10:FF:000003 | Leucine--tRNA ligase | - | - | 638 | 810 | 2.3E-79 |
| FunFam | G3DSA:3.40.50.620:FF:000124 | Leucine--tRNA ligase | - | - | 365 | 583 | 1.1E-100 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.