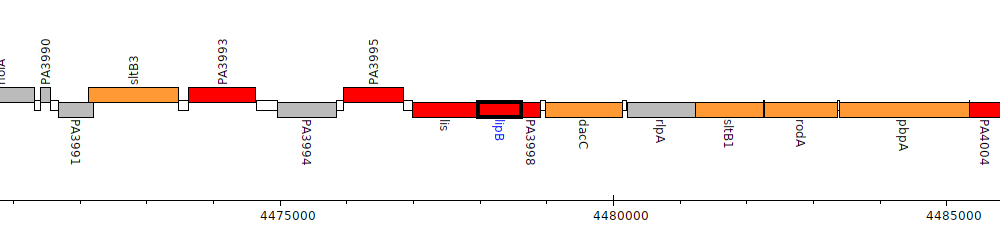
Pseudomonas aeruginosa PAO1, PA3997 (lipB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006554 | lysine catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006744 | ubiquinone biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0046656 | folic acid biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004619 | phosphoglycerate mutase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0051188 | obsolete cofactor biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009249 | protein lipoylation |
Inferred from Sequence Model
Term mapped from: InterPro:cd16444
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0033819 | lipoyl(octanoyl) transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd16444
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Folate biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Lysine degradation |
ECO:0000037
not_recorded |
|||
| KEGG | pae00785 | Lipoic acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Ubiquinone biosynthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55681 | Class II aaRS and biotin synthetases | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 6 | 204 | 2.57E-66 |
| NCBIfam | TIGR00214 | JCVI: lipoyl(octanoyl) transferase LipB | IPR000544 | Octanoyltransferase | 28 | 194 | 4.6E-62 |
| PIRSF | PIRSF016262 | LPLase | IPR000544 | Octanoyltransferase | 4 | 215 | 1.7E-57 |
| Hamap | MF_00013 | Octanoyltransferase [lipB]. | IPR000544 | Octanoyltransferase | 4 | 205 | 37.337898 |
| Gene3D | G3DSA:3.30.930.10 | Bira Bifunctional Protein; Domain 2 | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 3 | 202 | 5.7E-62 |
| FunFam | G3DSA:3.30.930.10:FF:000020 | Octanoyltransferase | - | - | 7 | 202 | 8.6E-92 |
| PANTHER | PTHR10993 | OCTANOYLTRANSFERASE | - | - | 8 | 203 | 1.3E-47 |
| CDD | cd16444 | LipB | IPR000544 | Octanoyltransferase | 7 | 202 | 2.02555E-111 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.