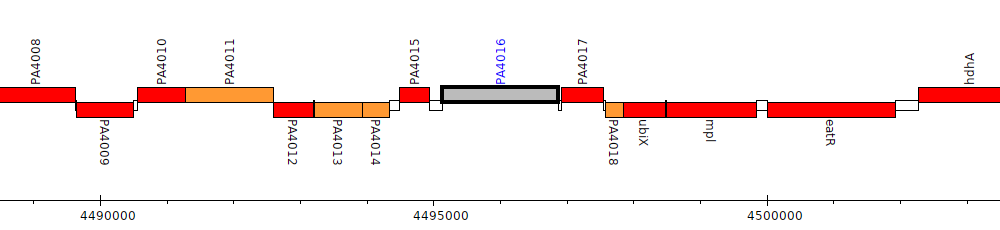
Pseudomonas aeruginosa PAO1, PA4016
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PF01650
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008233 | peptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01650
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 175 | 196 | 3.1 |
| Gene3D | G3DSA:3.40.50.1460 | - | - | - | 388 | 544 | 1.5E-13 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 63 | 85 | 2.3E-4 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 129 | 150 | 6.6 |
| Pfam | PF01650 | Peptidase C13 family | IPR001096 | Peptidase C13, legumain | 342 | 547 | 1.3E-43 |
| Coils | Coil | Coil | - | - | 506 | 526 | - |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 109 | 124 | 0.94 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 38 | 59 | 4.0 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 84 | 105 | 0.56 |
| SUPERFAMILY | SSF82185 | Histone H3 K4-specific methyltransferase SET7/9 N-terminal domain | - | - | 127 | 248 | 4.71E-25 |
| Gene3D | G3DSA:2.20.110.10 | - | - | - | 156 | 205 | 3.0E-10 |
| SUPERFAMILY | SSF52129 | Caspase-like | IPR029030 | Caspase-like domain superfamily | 392 | 530 | 6.45E-6 |
| Gene3D | G3DSA:2.20.110.10 | - | - | - | 206 | 252 | 1.2E-7 |
| PANTHER | PTHR46614 | MORN REPEAT-CONTAINING PROTEIN 4 | - | - | 35 | 123 | 1.4E-58 |
| SUPERFAMILY | SSF82185 | Histone H3 K4-specific methyltransferase SET7/9 N-terminal domain | - | - | 35 | 122 | 1.57E-20 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 154 | 175 | 0.083 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 107 | 128 | 1.5 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 61 | 82 | 13.0 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 177 | 199 | 3.0E-4 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 152 | 173 | 6.6 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 198 | 219 | 4.1E-5 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 267 | 288 | 8.0 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 131 | 153 | 0.0078 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 224 | 245 | 0.34 |
| SMART | SM00698 | morn | IPR003409 | MORN repeat | 244 | 265 | 21.0 |
| Gene3D | G3DSA:2.20.110.10 | - | - | - | 65 | 129 | 9.4E-8 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 86 | 108 | 0.13 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 269 | 291 | 0.041 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 163 | 197 | - |
| Gene3D | G3DSA:2.20.110.10 | - | - | - | 253 | 312 | 4.6E-6 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 166 | 182 | - |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 200 | 222 | 7.2E-6 |
| Pfam | PF02493 | MORN repeat | IPR003409 | MORN repeat | 40 | 60 | 0.042 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.