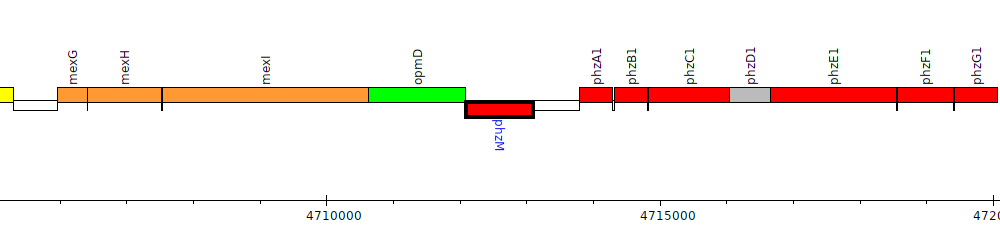
Pseudomonas aeruginosa PAO1, PA4209 (phzM)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
17253782 | Reviewed by curator |
| Molecular Function | GO:0003824 | catalytic activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0002047 | phenazine biosynthetic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0008171 | O-methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43712
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53335
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Phenazine biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY-6666 | pyocyanin biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | PWYB-5652 | Phenazine biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43712 | PUTATIVE (AFU_ORTHOLOGUE AFUA_4G14580)-RELATED | IPR016461 | O-methyltransferase COMT-type | 12 | 330 | 5.7E-47 |
| Pfam | PF00891 | O-methyltransferase domain | IPR001077 | O-methyltransferase domain | 115 | 314 | 2.3E-28 |
| Pfam | PF16864 | Dimerisation domain | IPR031725 | Acetylserotonin O-methyltransferase, dimerisation domain | 12 | 86 | 1.5E-6 |
| SUPERFAMILY | SSF53335 | S-adenosyl-L-methionine-dependent methyltransferases | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 95 | 330 | 1.03E-39 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 1 | 88 | 1.1E-22 |
| SUPERFAMILY | SSF46785 | Winged helix DNA-binding domain | IPR036390 | Winged helix DNA-binding domain superfamily | 10 | 89 | 5.99E-14 |
| PIRSF | PIRSF005739 | O-mtase | IPR016461 | O-methyltransferase COMT-type | 1 | 332 | 1.7E-37 |
| Gene3D | G3DSA:3.40.50.150 | Vaccinia Virus protein VP39 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase superfamily | 89 | 334 | 2.7E-66 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.