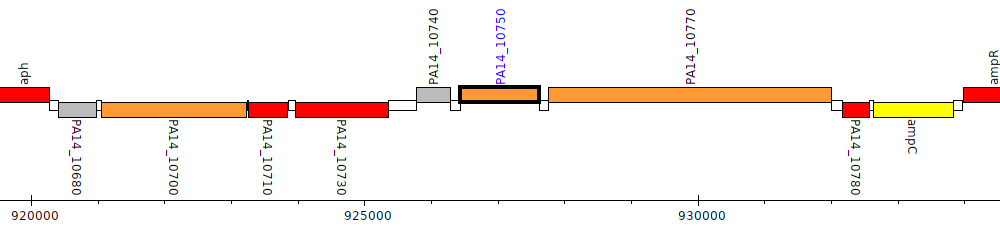
Pseudomonas aeruginosa UCBPP-PA14, PA14_10750
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 9 | 384 | 9.4E-40 |
| SUPERFAMILY | SSF103473 | MFS general substrate transporter | IPR036259 | MFS transporter superfamily | 7 | 385 | 1.7E-54 |
| Hamap | MF_00517 | Probable sugar efflux transporter [sotB]. | IPR023495 | Sugar efflux transporter, putative | 1 | 390 | 29.900505 |
| PANTHER | PTHR43124 | PURINE EFFLUX PUMP PBUE | - | - | 11 | 392 | 5.3E-97 |
| CDD | cd17324 | MFS_NepI_like | - | - | 15 | 366 | 2.83247E-80 |
| Pfam | PF07690 | Major Facilitator Superfamily | IPR011701 | Major facilitator superfamily | 18 | 348 | 2.1E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.