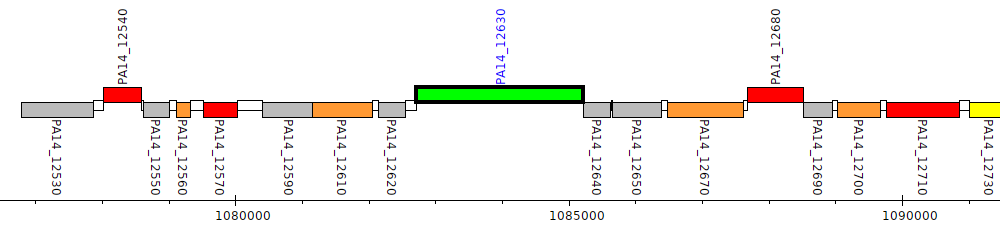
Pseudomonas aeruginosa UCBPP-PA14, PA14_12630
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004386 | helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04408
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00847 | ha2_5 | IPR007502 | Helicase-associated domain | 385 | 488 | 7.4E-17 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 67 | 154 | 1.94E-9 |
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 202 | 325 | 3.0E-14 |
| Gene3D | G3DSA:1.20.120.1080 | - | - | - | 370 | 484 | 3.0E-13 |
| Pfam | PF04408 | Helicase associated domain (HA2) | IPR007502 | Helicase-associated domain | 386 | 446 | 5.7E-8 |
| NCBIfam | TIGR01970 | JCVI: ATP-dependent helicase HrpB | IPR010225 | ATP-dependent helicase HrpB | 67 | 829 | 0.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 33 | 60 | - |
| FunFam | G3DSA:1.20.120.1080:FF:000055 | ATP-dependent helicase HrpB | - | - | 369 | 481 | 1.9E-34 |
| PANTHER | PTHR43519 | ATP-DEPENDENT RNA HELICASE HRPB | - | - | 67 | 829 | 0.0 |
| CDD | cd18791 | SF2_C_RHA | - | - | 176 | 334 | 1.62327E-64 |
| SMART | SM00490 | helicmild6 | IPR001650 | Helicase, C-terminal | 222 | 326 | 6.5E-19 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 39 | 174 | 1.5E-30 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 175 | 343 | 9.7E-56 |
| PIRSF | PIRSF005496 | ATP_hel_HrpB | IPR010225 | ATP-dependent helicase HrpB | 64 | 830 | 0.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 35 | 49 | - |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 173 | 438 | 6.02E-50 |
| Pfam | PF08482 | ATP-dependent helicase C-terminal | IPR013689 | ATP-dependent RNA helicase HrpB, C-terminal | 686 | 817 | 2.4E-52 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.