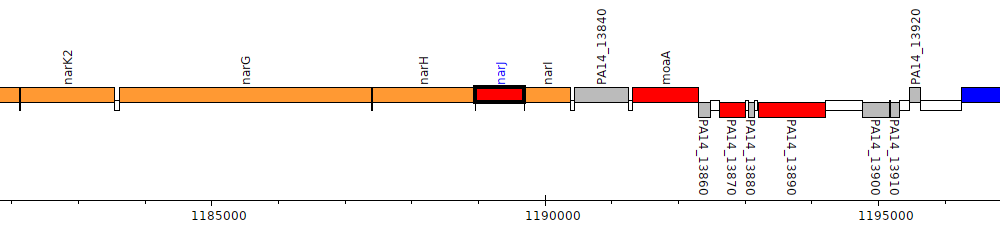
Pseudomonas aeruginosa UCBPP-PA14, PA14_13810 (narJ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051082 | unfolded protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00684
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0051131 | chaperone-mediated protein complex assembly |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00684
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 201 | 227 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 207 | 227 | - |
| PANTHER | PTHR43680 | NITRATE REDUCTASE MOLYBDENUM COFACTOR ASSEMBLY CHAPERONE | IPR003765 | Nitrate reductase chaperone, NarJ | 1 | 245 | 2.9E-70 |
| Gene3D | G3DSA:1.10.3480.10 | - | - | - | 1 | 168 | 3.4E-8 |
| Pfam | PF02613 | Nitrate reductase delta subunit | IPR020945 | DMSO/Nitrate reductase chaperone | 40 | 160 | 5.2E-10 |
| SUPERFAMILY | SSF89155 | TorD-like | IPR036411 | TorD-like superfamily | 6 | 166 | 6.41E-30 |
| NCBIfam | TIGR00684 | JCVI: nitrate reductase molybdenum cofactor assembly chaperone | IPR003765 | Nitrate reductase chaperone, NarJ | 13 | 164 | 1.2E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.