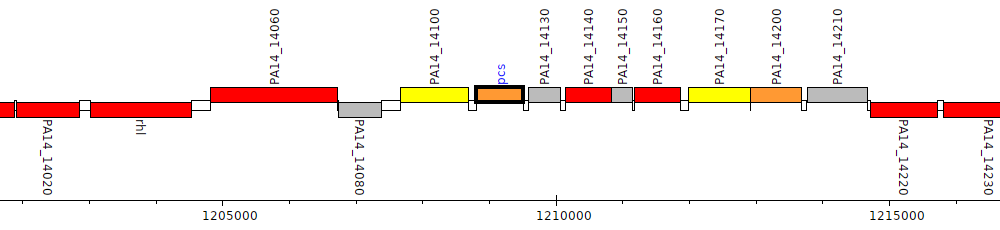
Pseudomonas aeruginosa UCBPP-PA14, PA14_14110 (pcs)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050520 | phosphatidylcholine synthase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3857
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
12169604 | |
| Molecular Function | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008654 | phospholipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00564 | Glycerophospholipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.120.1760 | - | IPR043130 | CDP-alcohol phosphatidyltransferase, transmembrane domain | 15 | 180 | 7.1E-14 |
| FunFam | G3DSA:1.20.120.1760:FF:000034 | Phosphatidylcholine synthase | - | - | 15 | 179 | 1.6E-110 |
| PIRSF | PIRSF000851 | PcS | IPR026027 | Phosphatidylcholine synthase Pcs | 2 | 238 | 1.1E-89 |
| Pfam | PF01066 | CDP-alcohol phosphatidyltransferase | IPR000462 | CDP-alcohol phosphatidyltransferase | 21 | 163 | 1.3E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.