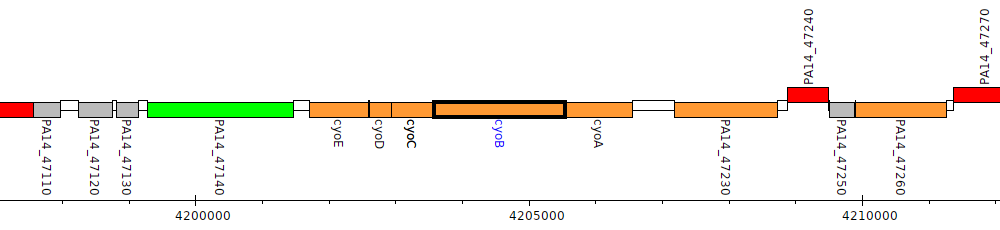
Pseudomonas aeruginosa UCBPP-PA14, PA14_47190 (cyoB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009060 | aerobic respiration |
Inferred from Sequence Model
Term mapped from: InterPro:PF00115
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00115
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00115
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF00115
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016682 | oxidoreductase activity, acting on diphenols and related substances as donors, oxygen as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02843
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00190 | Oxidative phosphorylation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF81442 | Cytochrome c oxidase subunit I-like | IPR036927 | Cytochrome c oxidase-like, subunit I superfamily | 43 | 559 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 47 | 72 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 411 | 430 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 97 | 120 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 124 | 148 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 198 | 216 | 1.5E-95 |
| FunFam | G3DSA:1.20.210.10:FF:000002 | Cytochrome o ubiquinol oxidase, subunit I | - | - | 52 | 552 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 166 | 178 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 461 | 482 | 1.5E-95 |
| Gene3D | G3DSA:1.20.210.10 | - | IPR036927 | Cytochrome c oxidase-like, subunit I superfamily | 31 | 564 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 383 | 401 | 1.5E-95 |
| NCBIfam | TIGR02843 | JCVI: cytochrome o ubiquinol oxidase subunit I | IPR014207 | Cytochrome o ubiquinol oxidase, subunit I | 2 | 647 | 0.0 |
| CDD | cd01662 | Ubiquinol_Oxidase_I | - | - | 48 | 552 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 348 | 369 | 1.5E-95 |
| PANTHER | PTHR10422 | CYTOCHROME C OXIDASE SUBUNIT 1 | IPR000883 | Cytochrome c oxidase subunit I | 45 | 564 | 0.0 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 227 | 246 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 278 | 299 | 1.5E-95 |
| PRINTS | PR01165 | Cytochrome c oxidase subunit I signature | IPR000883 | Cytochrome c oxidase subunit I | 324 | 339 | 1.5E-95 |
| Pfam | PF00115 | Cytochrome C and Quinol oxidase polypeptide I | IPR000883 | Cytochrome c oxidase subunit I | 56 | 504 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.