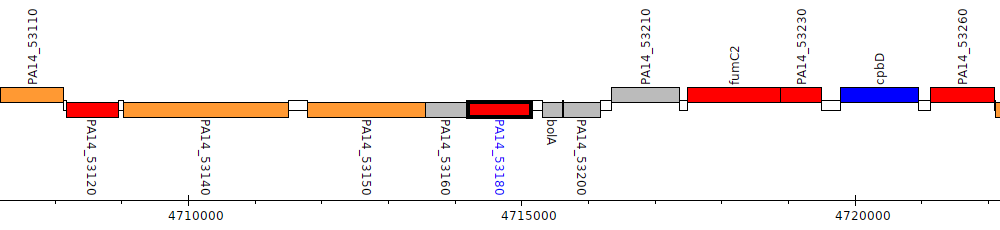
Pseudomonas aeruginosa UCBPP-PA14, PA14_53180
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00581 | Rhodanese-like domain | IPR001763 | Rhodanese-like domain | 115 | 210 | 5.9E-7 |
| Gene3D | G3DSA:3.40.250.10 | - | IPR036873 | Rhodanese-like domain superfamily | 97 | 251 | 2.4E-52 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 1 | 96 | 1.6E-29 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 282 | 300 | - |
| SMART | SM00450 | rhod_4 | IPR001763 | Rhodanese-like domain | 113 | 214 | 1.9E-15 |
| CDD | cd01518 | RHOD_YceA | - | - | 110 | 210 | 5.27464E-55 |
| Pfam | PF17773 | UPF0176 acylphosphatase like domain | IPR040503 | tRNA uridine(34) hydroxylase, N-terminal | 5 | 95 | 3.5E-25 |
| PANTHER | PTHR43268 | THIOSULFATE SULFURTRANSFERASE/RHODANESE-LIKE DOMAIN-CONTAINING PROTEIN 2 | IPR020936 | tRNA uridine(34) hydroxylase | 3 | 280 | 1.9E-90 |
| SUPERFAMILY | SSF52821 | Rhodanese/Cell cycle control phosphatase | IPR036873 | Rhodanese-like domain superfamily | 90 | 225 | 1.37E-28 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 282 | 312 | - |
| Hamap | MF_00469 | tRNA uridine(34) hydroxylase [trhO]. | IPR020936 | tRNA uridine(34) hydroxylase | 2 | 254 | 46.232983 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.