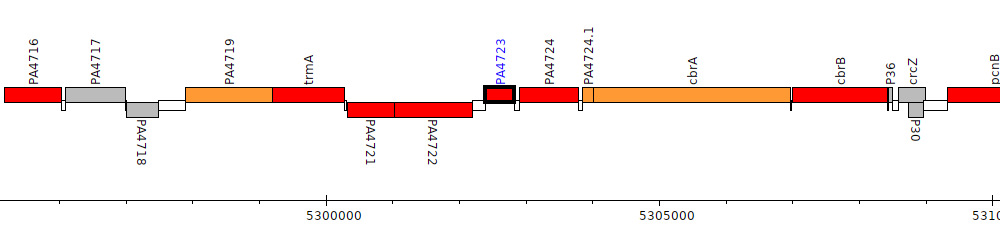
Pseudomonas aeruginosa PAO1, PA4723
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0051055 | negative regulation of lipid biosynthetic process | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
11160083 | Reviewed by curator |
| Biological Process | GO:2000679 | positive regulation of transcription regulatory region DNA binding | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15853892 | Reviewed by curator |
| Biological Process | GO:0050896 | response to stimulus | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006979 | response to oxidative stress | Inferred from Genetic Interaction | ECO:0006091 |
35729352 | Reviewed by curator |
| Molecular Function | GO:0070063 | RNA polymerase binding | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
21255113 | Reviewed by curator |
| Biological Process | GO:1901857 | positive regulation of cellular respiration | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
18203836 | Reviewed by curator |
| Biological Process | GO:0045892 | negative regulation of transcription, DNA-templated | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15853892 | Reviewed by curator |
| Biological Process | GO:0045861 | negative regulation of proteolysis | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
11160083 | Reviewed by curator |
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01258
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR33823 | RNA POLYMERASE-BINDING TRANSCRIPTION FACTOR DKSA-RELATED | - | - | 29 | 145 | 2.9E-35 |
| PRINTS | PR00618 | DksA/TraR zinc finger signature | IPR020460 | Zinc finger, DksA/TraR C4-type, bacteria | 111 | 122 | 7.6E-5 |
| SUPERFAMILY | SSF109635 | DnaK suppressor protein DksA, alpha-hairpin domain | IPR037187 | DksA, N-terminal domain superfamily | 11 | 106 | 6.8E-34 |
| NCBIfam | TIGR02420 | JCVI: RNA polymerase-binding protein DksA | IPR012784 | RNA polymerase-binding transcription factor DksA | 29 | 137 | 2.6E-51 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 23 | - |
| Gene3D | G3DSA:1.20.120.910 | - | - | - | 2 | 148 | 5.2E-42 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 18 | - |
| Pfam | PF01258 | Prokaryotic dksA/traR C4-type zinc finger | IPR000962 | Zinc finger, DksA/TraR C4-type | 108 | 140 | 8.4E-11 |
| Hamap | MF_00926 | RNA polymerase-binding transcription factor DksA [dksA]. | IPR012784 | RNA polymerase-binding transcription factor DksA | 25 | 147 | 31.0874 |
| SUPERFAMILY | SSF57716 | Glucocorticoid receptor-like (DNA-binding domain) | - | - | 108 | 145 | 2.71E-10 |
| PRINTS | PR00618 | DksA/TraR zinc finger signature | IPR020460 | Zinc finger, DksA/TraR C4-type, bacteria | 123 | 135 | 7.6E-5 |
| FunFam | G3DSA:1.20.120.910:FF:000001 | RNA polymerase-binding transcription factor DksA | - | - | 2 | 148 | 1.6E-80 |
| Coils | Coil | Coil | - | - | 86 | 106 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.