Pseudomonas aeruginosa PAO1, PA1254 (lhpC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
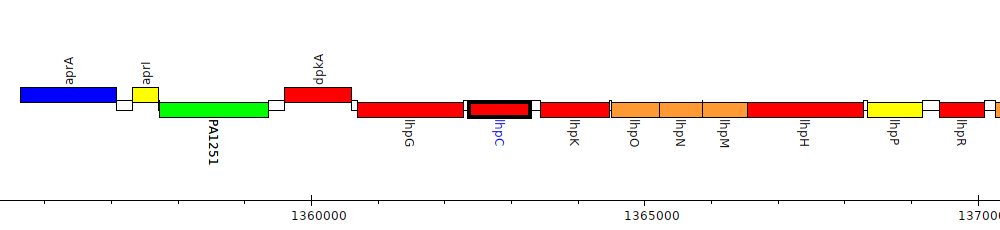
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA1254
|
| Name |
lhpC
|
| Replicon | chromosome |
| Genomic location | 1362371 - 1363288 (- strand) |
| Transposon Mutants | 1 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 5 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_249945.1 |
| GI | 15596451 |
| Affymetrix | PA1254_at |
| DNASU | PaCD00006579 |
| Entrez | 881320 |
| GenBank | AAG04643.1 |
| INSDC | AAG04643.1 |
| NCBI Locus Tag | PA1254 |
| protein_id(GenBank) | gb|AAG04643.1|AE004555_3|gnl|PseudoCAP|PA1254 |
| TIGR | NTL03PA01255 |
| UniParc | UPI00000C52D0 |
| UniProtKB Acc | Q9I490 |
| UniProtKB ID | Q9I490_PSEAE |
| UniRef100 | UniRef100_A0A0C7D604 |
| UniRef50 | UniRef50_U1LTR1 |
| UniRef90 | UniRef90_A0A022NV25 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
delta1-pyrroline-4-hydroxy-2-carboxylate deaminase, LphC
|
| Synonyms |
probable dihydrodipicolinate synthetase |
| Evidence for Translation | |
| Charge (pH 7) | -5.73 |
| Kyte-Doolittle Hydrophobicity Value | 0.107 |
| Molecular Weight (kDa) | 32.9 |
| Isoelectric Point (pI) | 5.41 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 632 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG000222 (1144 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA1254
Search term: lhpC
|
Human Homologs
References
|
Molecular characterization of LhpR in control of hydroxyproline catabolism and transport in Pseudomonas aeruginosa PAO1.
Li G, Lu CD
Microbiology (Reading) 2016 Jul;162(7):1232-1242
PubMed ID: 27145750
|
|
Alpha-ketoglutaric semialdehyde dehydrogenase of Pseudomonas. Properties of the purified enzyme induced by hydroxyproline and of the glucarate-induced and constitutive enzymes.
Adams E, Rosso G
J Biol Chem 1967 Apr 25;242(8):1802-14
PubMed ID: 6024771
|
|
Identification and characterization of D-hydroxyproline dehydrogenase and Delta1-pyrroline-4-hydroxy-2-carboxylate deaminase involved in novel L-hydroxyproline metabolism of bacteria: metabolic convergent evolution.
Watanabe S, Morimoto D, Fukumori F, Shinomiya H, Nishiwaki H, Kawano-Kawada M, Sasai Y, Tozawa Y, Watanabe Y
J Biol Chem 2012 Sep 21;287(39):32674-88
PubMed ID: 22833679
|