Pseudomonas aeruginosa PAO1, PA4856 (retS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
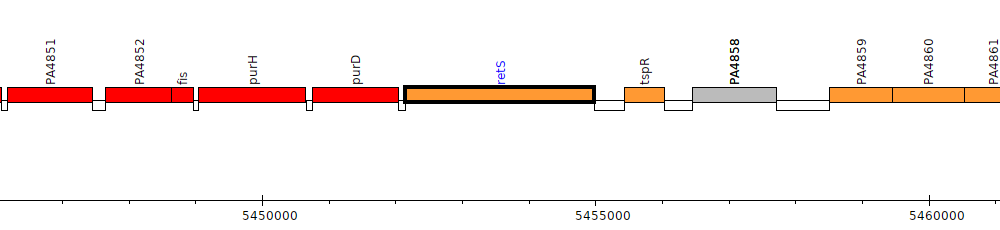
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa PAO1 (Stover et al., 2000)
GCF_000006765.1|latest |
| Locus Tag |
PA4856
|
| Name |
retS
Synonym: rtsM |
| Replicon | chromosome |
| Genomic location | 5452150 - 5454978 (+ strand) |
| Transposon Mutants | 5 transposon mutants in PAO1 |
| Transposon Mutants in orthologs | 1 transposon mutants in orthologs |
Cross-References
| RefSeq | NP_253543.1 |
| GI | 15600049 |
| Affymetrix | PA4856_at |
| Entrez | 877881 |
| GenBank | AAG08241.1 |
| INSDC | AAG08241.1 |
| NCBI Locus Tag | PA4856 |
| protein_id(GenBank) | gb|AAG08241.1|AE004899_2|gnl|PseudoCAP|PA4856 |
| TIGR | NTL03PA04857 |
| UniParc | UPI00000C5E3E |
| UniProtKB Acc | Q9HUV7 |
| UniProtKB ID | Q9HUV7_PSEAE |
| UniRef100 | UniRef100_Q9HUV7 |
| UniRef50 | UniRef50_Q4KIX7 |
| UniRef90 | UniRef90_Q9HUV7 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
RetS (Regulator of Exopolysaccharide and Type III Secretion)
|
| Synonyms |
RtsM |
| Evidence for Translation |
Identified using nanoflow high-pressure liquid chromatography (HPLC) in conjunction with microelectrospray ionization on LTQ XL mass spectrometer (PMID:24291602).
|
| Charge (pH 7) | -5.05 |
| Kyte-Doolittle Hydrophobicity Value | 0.111 |
| Molecular Weight (kDa) | 103.8 |
| Isoelectric Point (pI) | 6.47 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 3JYB | TRANSFERASE | 09/21/09 | Crystal Structure of the RetS periplasmic domain | Pseudomonas aeruginosa | 2.04 | X-RAY DIFFRACTION | 100.0 |
| 6DK8 | SIGNALING PROTEIN | 05/29/18 | RetS kinase region without cobalt | Pseudomonas aeruginosa | 3.8 | X-RAY DIFFRACTION | 100.0 |
| 2XBZ | TRANSFERASE | 04/15/10 | Multiple oligomeric forms of the Pseudomonas aeruginosa RetS sensor domain modulate accessibility to the ligand-binding site | PSEUDOMONAS AERUGINOSA | 2.65 | X-RAY DIFFRACTION | 100.0 |
| 7N0E | SIGNALING PROTEIN | 05/25/21 | Co-complex of the histidine kinase region of RetS and the dimerization and histidine phosphotransfer domain of GacS | Pseudomonas aeruginosa | 2.3 | X-RAY DIFFRACTION | 100.0 |
| 6DK7 | SIGNALING PROTEIN | 05/29/18 | RetS histidine kinase region with cobalt | Pseudomonas aeruginosa | 2.6 | X-RAY DIFFRACTION | 100.0 |
Virulence Evidence
| Source 1 | |
| Database | PseudoCAP |
| Category | |
| Evidence |
ECO:0000314
direct assay evidence used in manual assertion
|
| Host Organism | Homo sapiens |
| Notes | |
| Infection model | Lactate dehydrogenase (LDH) assay of infected HeLa cells |
| Reference | 15522089 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 618 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG004485 (533 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PA4856
Search term: retS
|
Human Homologs
References
|
A novel sensor kinase-response regulator hybrid regulates type III secretion and is required for virulence in Pseudomonas aeruginosa.
Laskowski MA, Osborn E, Kazmierczak BI
Mol Microbiol 2004 Nov;54(4):1090-103
PubMed ID: 15522089
|
|
A signaling network reciprocally regulates genes associated with acute infection and chronic persistence in Pseudomonas aeruginosa.
Goodman AL, Kulasekara B, Rietsch A, Boyd D, Smith RS, Lory S
Dev Cell 2004 Nov;7(5):745-54
PubMed ID: 15525535
|
|
The genetic basis for the commitment to chronic versus acute infection in Pseudomonas aeruginosa.
Yahr TL, Greenberg EP
Mol Cell 2004 Nov 19;16(4):497-8
PubMed ID: 15546607
|
|
Genome-wide identification of Pseudomonas aeruginosa exported proteins using a consensus computational strategy combined with a laboratory-based PhoA fusion screen.
Lewenza S, Gardy JL, Brinkman FS, Hancock RE
Genome Res 2005 Feb;15(2):321-9
PubMed ID: 15687295
|
|
The lemA gene required for pathogenicity of Pseudomonas syringae pv. syringae on bean is a member of a family of two-component regulators.
Hrabak EM, Willis DK
J Bacteriol 1992 May;174(9):3011-20
PubMed ID: 1314807
|